DAVID Documentation: Ortholog
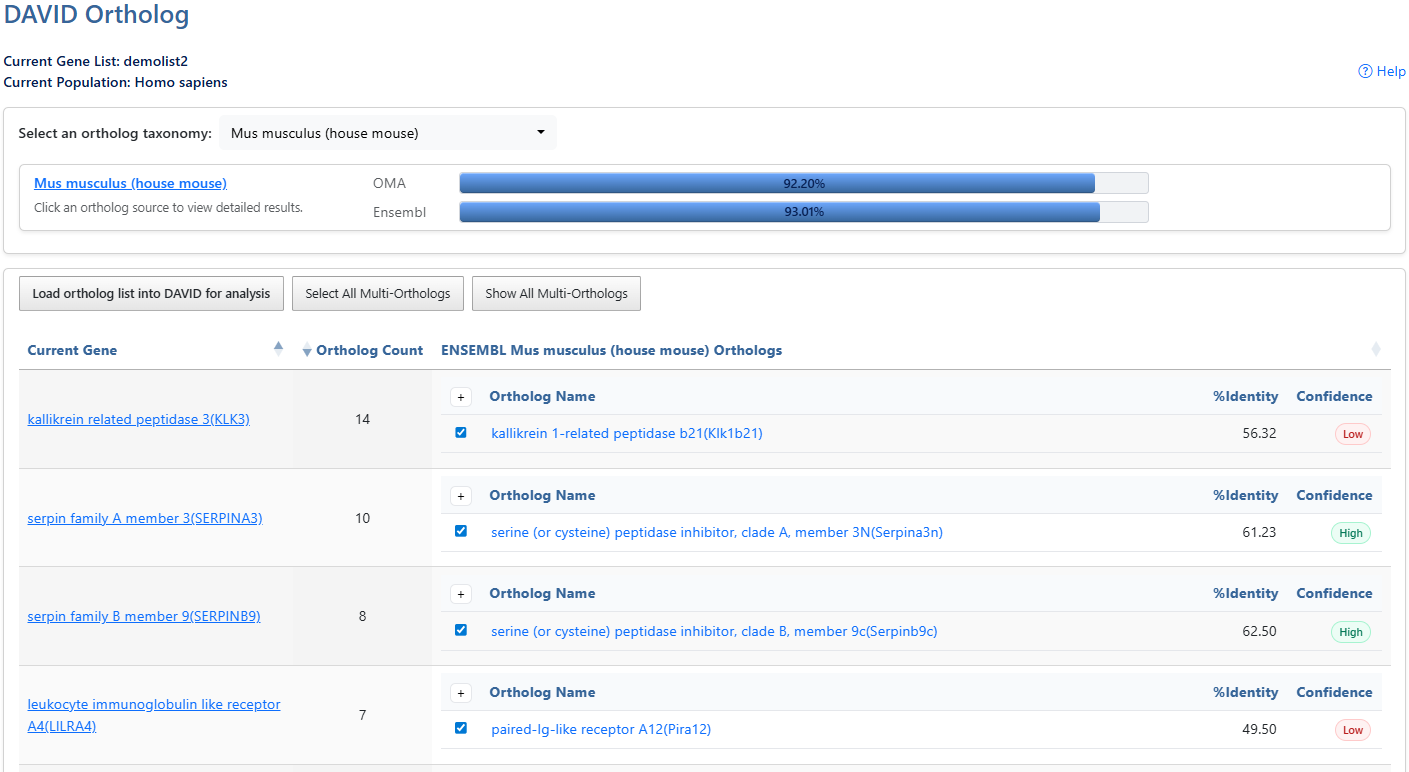
DAVID Ortholog enables cross-species functional analysis by converting a source gene list into an orthologous list in a user-selected target species. Ortholog mapping data from Orthologous MAtrix (OMA) and Ensembl Compara are integrated into the DAVID Knowledgebase through DAVID Gene identifiers,
-
Current Gene List & PopulationDisplays the active gene list and selected species or background population.
-
Help and Switch to Classic VersionLink to this help page and provides access to the legacy Gene List Report interface.
-
Ortholog TaxonomyClick the dropdown box to select a target ortholog taxonomy (species). After clicking on the box, a search box is also available to type and search for taxonomies. Click on the taxonomy to select, then the ortholog sources will populate.
-
SpeciesClick the Species link to go to NCBI taxonomy entry.
-
OMAShows the percentage of genes from the list with an ortholog available for the selected taxonomy from Orthologous Matrix (OMA) Click the percentage bar to retrieve orthologs from that source.
-
EnsemblShows the percentage of genes from the list with an ortholog available for the selected taxonomy from Ensembl Compara. Click the percentage bar to retrieve orthologs from that source.
-
Load to DAVIDLoad selected orthologs into a DAVID list for downstream analysis.
-
Select AllSelect All Multi-Orthologs will select all available orthologs. Click again to Deselect all. Click again to select only the first ortholog if there are multiple available (default).
-
ShowShow/Hide all multi-orthologs
-
Sortable ColumnsClick any column header to sort by User ID, Expression Value, DAVID Gene Name, Related Genes (RG), or Species.
-
Gene nameClick the gene name link to explore the full gene report. For more information, see the Gene Report documentation page.
-
Ortholog countNumber of ortholog genes available for the user gene in the selected taxonomy.
-
ExpandShow/Hide all multi-orthologs
-
SelectSelected orthologs are loaded as a new DAVID list when clicking the Load ortholog list into DAVID for analysis button.
-
Ortholog Gene nameClick the gene name link to explore the full gene report. For more information, see the Gene Report documentation page.
-
IdentitySequence identity between user gene and ortholog gene. Higher meaning more identical.
-
ConfidenceConfidence score. Only available for Ensembl results.